Relaxation Assays
The relaxation of negative or positive supercoiled DNA is catalysed by both type I and type II DNA topoisomerases, however the process by which they catalyse these reactions differs depending on the type. The process in the type II topoisomerases is ATP-dependent and occurs through the cleavage and religation of the double-stranded DNA in steps of 2. While it is independent of ATP with type I topoisomerases where only one strand of the DNA is cleaved.
For an overview of the reaction please see: , , & (2021). DNA topoisomerases: Advances in understanding of cellular roles and multi-protein complexes via structure-function analysis. BioEssays, 43, e2000286.
A typical reaction is as follows:
1 U of enzyme is incubated with 0.5 µg of supercoiled pBR322 DNA in a 30 µl reaction at 37°C for 30 minutes in 1X Assay Buffer
Each reaction is stopped by the addition of 30 µl chloroform/iso-amyl alcohol (24/1) and 30 µl Stop Buffer (STEB: 40% sucrose, 100 mM Tris.HCl ( pH 7.5), 100 mM EDTA, 0.5 mg/ml bromophenol blue), before being loaded on an agarose gel. (0.8%-1.0%: w/v)
Gels can be run either in TBE (90 mM Tris.Borate, 2 mM EDTA) or TAE (40 mM Tris.Acetate, 2 mM EDTA) although the resolution of supercoiled pBR322 is much better if run in TAE at a maximum of 90V for 2 - 3 hours.
Typical Gels:
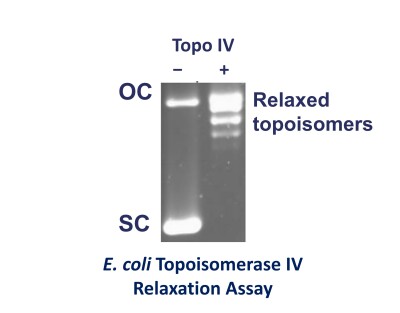
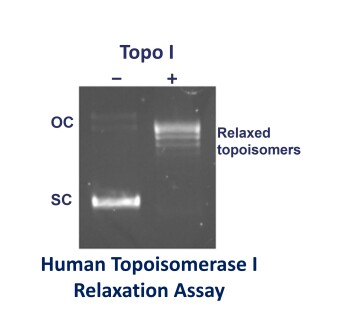
The substrate used usually consists of two bands on a gel. The upper one is open-circular (nicked) DNA and the faster migrating band is negatively supercoiled (closed circular) plasmid.
Treatment of negatively supercoiled pBR322 with a topo I, topo II or topo IV converts the supercoiled form to the relaxed form and this consists of a series of bands on a gel which are different topoisomers (DNAs of different linking number).
Intercalators
Problems can arise in assays if there are contaminants such as ethidium bromide or Chloroquine in the gel tanks. These chemicals are intercalators and can affect the mobility of the DNA leading to confusing results. If there is an intercalator present in the buffer the relaxed topoisomers have increased mobility and run in roughly the same place as negatively supercoiled plasmid. Depending on the amount of intercalator, the negatively supercoiled DNA may have slightly less mobility than normal.
Nucleases
If there are contaminating nucleases present in the assay (due to contaminated assay buffers or impure fractions from cell expression) there may be a significant increase in nicking leading to an increase in the amount of open-circular DNA and possibly the production of linear DNA. This runs as a single band between the OC and SC.
ATP
A loss of activity with previously active enzyme may be due to deterioration of the ATP in the buffer, although, the 5X Assay Buffer has been developed to minimize this. If loss of activity occurs, the addition of extra ATP to the reaction will overcome the problem.
