Investigation of Protein-DNA Interactions with ReDCaT Chip
Background
The interaction of specific DNA sequences with proteins is central to many cellular processes including transcription, replication, repair, and gene regulation. The recognition sequence can be determined by a number of techniques, such as electrophoretic mobility shift assays (EMSA) and filter binding assays.
The technique of Surface Plasmon Resonance (SPR) is ideally suited as an alternative technique to measure these interactions because it is quantitative, simple to perform, reproducible, can be automated, and requires very little sample. RedcaT (Reusable DNA Capture Technology) uses SPR but allows a single chip to be reused multiple times so that many DNA sequences can be tested at low cost.
How It Works
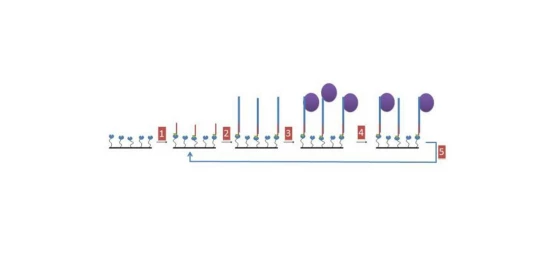
- A biotin-labelled, single-stranded oligo is immobilised onto a streptavidin-coated chip.
- Two oligos are annealed together to produce a double-stranded oligo of the sequence to be tested and a single-stranded overhang which is complementary to the immobilised sequence. This is then immobilized via this single-stranded sequence to the biotin-immobilised oligo.
- After washing away unbound oligo, the protein of interest is passed over the immobilised double-stranded oligo. If protein is bound then a signal is produced.
- The complex can then be washed with different buffers which may remove weakly bound proteins.
- The chip is finally washed to remove the target oligo and any remaining bound protein, leaving the biotin-labelled oligo available for reuse for the next target sequence.
What Redcat Can Do
Determine if a protein can bind to a particular DNA sequence.
Examples would include:
- If you have a sequence which you believe is the binding site for your protein then this can be tested and compared with other sequences. For example, sequences from a MEME-predicted binding site.
- Identifying the binding sequence from a promoter region by breaking it into a number of overlapping sequences and comparing the interactions with your protein.
- Determine the footprint of a protein on a DNA sequence (e.g. to determine the minimum length of sequence needed for co-crystallisation trials) by testing sequences of varying length.
- Rank sequences in a regulon.
- Compare DNA sequences to identify key bases involved in an interaction. Compare wild-type and mutated DNA sequences.
- Identify key residues in a protein involved in the DNA interaction by comparing the binding of wild-type and mutant versions of the protein.
- Investigate the effect of inhibitors and effectors on the DNA-protein interaction.
Advantages
- Does not require labeling of protein or DNA
- Uses low amounts of protein compared with other techniques
- Fast
- Can detect interactions not always seen with EMSA because it does not rely on the protein-DNA interaction being resistant to the electrophoresis step
Pricing
Our prices for the Redcat service are currently under review. Please enquire for more information.
There is a setup fee which covers the cost of setting up the assay for a new study. So if we have tested your protein and you subsequently decide to have more oligos tested then there will be no second setup fee for these, just the sample fees. We will also store your protein for a period of time, if requested, so that if you do decide to have more DNA sequences tested then you don’t have to send more protein.
What is included in the price?
Controls
A control run is made at the start and end of each set of samples to ensure that the process is working correctly. This uses a protein and DNA sequence which give a known response.
The sample is also run against the control DNA.
Output of results
We provide a report containing the output from the interaction experiments and a brief analysis.
How to Order and Requirements
How much protein to send
This depends on the number of runs to be made but a minimum of 10μl at 100μM concentration.
How to send the protein.
The protein should be at a concentration of 100μM and sent on dry ice.
Protein quality
You should ensure that your protein is soluble in conditions similar to the assay buffer (0.01 M HEPES pH 7.4, 0.15 M NaCl, 3 mM EDTA, 0.05% v/v Surfactant P20). Your protein should also be filtered (0.22 μm filter). Please also send information about any small molecules included in the sample buffer (such as DMSO) as these will need to be incorporated into the running buffer.
Description of the DNA sequences to be tested. The sequence can be up to 40 bases in length (we can perform the experiment with longer sequences but these are not standard lengths so please contact us). For each double-stranded DNA sequence to be tested please insert the complementary sequences (forward and reverse) into the order from attached (Redcat order form). You don’t need to add the single-stranded section for immobilization, we will do this.
Alternatively, and we recommend that you do this, you can download a copy of the program to generate overlapping oligos (PooP) using the link below. This program enables you to input your sequence of interest and then generate a series of overlapping DNA fragments to which you can attach the ssDNA linker required for the ReDCaT method. You can specify how long you wish each fragment to be and how many base pairs should be incorporated into the overlap.
